 |
|
CAMPR3 (Collection of Anti-Microbial Peptides) has been created to expand and accelerate antimicrobial peptide family-based studies. Antimicrobial peptides have family-specific sequence composition which can be mined to discover and design novel AMPs. CAMPR3 contains information on the conserved sequence signatures captured as patterns and Hidden Markov Models (HMMs) in 1386 AMPs represented by 45 families. Information related to sequence, protein definition, accession numbers, activity, source organism, target organisms, protein family descriptions and links to databases like UniProt, PubMed and other antimicrobial peptide databases are provided for the benefit of users. The database also provides tools for sequence alignment, pattern creation, pattern and HMM-based search. Comprehensive documentation of the database can be obtained from the Help page. This work has been funded by grants from Department of Science & Technology, India and Indian Council of Medical Research. |
Sequences  |
Structures  |
Patents  |
Signatures 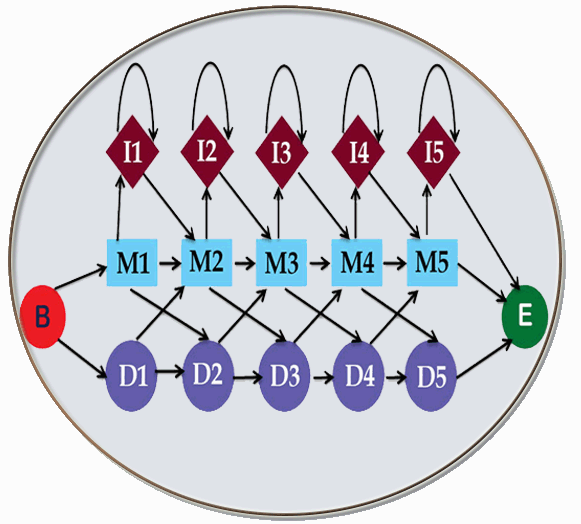 |
|
CAMPSign is a tool for identification of antimicrobial peptides belonging to one of the 45 major AMP families present in CAMPR3. For using CAMPSign Please Click here.... |
Citation :
Previous Citations :
|
| © Biomedical Informatics Centre, NIRRH, Mumbai |